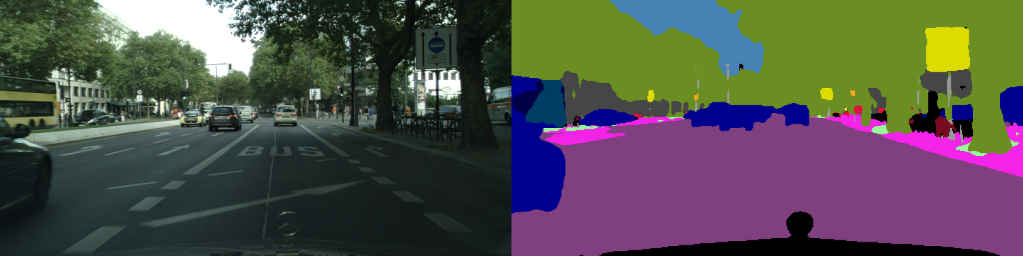
Introduction
Apache Airflow is a well known product for creating automated ETL (Extract-transform-load) pipelines to process raw data for training machine learning models. Similarly, Docker Swarm and Kubernetes are popular as management systems of containerized applications with multi-node/multi-cluster architectures.
In this post, we make use of these products together to create a model hyperparameter optimization pipeline that is easily scalable across many machines with one or more GPUs for accelerated computation. We need the system to have the following features:
- Ability to create a workflow graph so that the pipeline task dependencies can be clearly seen and controlled
- Ability to easily add and remove nodes (with GPU devices) before/during training
- Have pre-training tasks that have functions like uploading raw data into an annotation tool, waiting for labeling to finish, transforming annotated data in the format suitable for consumption by our model trainer etc
- Have a set of training and evaluation tasks that can train with different hyperparameters and different shuffled dataset sets
- Have post-training tasks that carry out functions like determining the best model based on calculated test metrics and uploading as well as versioning the weights file of this model to production
- Ability to monitor and report the status of each of the above tasks
This post uses the following versions of software:
- Apache Airflow – 2.2.4
- Docker – 20.10.16
- Kubernetes – 1.24.0
Additionally, we are referring to NVIDIA GPUs when GPUs are mentioned.
Designing the pipeline
Setting up
We can start designing our pipeline with Apache Airflow. To setup Airflow, we basically have two options – either install it locally or use docker images from DockerHub. If we intend to use Kubernetes for running and maintaining our pipeline, the former is advisable. On the other hand, if we want to stick with single machine or limited node architecture, the latter is better suited. Upgrading Airflow image version and avoiding dependency conflicts is much easier with docker images but then we will have to deal with docker-in-docker problems, i.e. when the Airflow inside the docker wants to control containers/nodes management tool (like Kubernetes and Docker) running on the host machine.
DAGs for pipeline
We can group all our tasks into three different cliques – which we will refer as DAGs (directed acyclic graph). Grouping tasks into DAGs not only makes the whole pipeline management much easier but also allows us to skip a set of tasks if we didn’t want them to run. For instance, if we already had labeled data, we would want to skip all the tasks responsible for labeling raw data. This would be much easier to do if we grouped labeling data into one DAG and similarly one each for others as appropriate.

The first one will be called Prelabeling DAG for handling data labeling, the second Training DAG will be responsible for finding the best set of hyperparameters and, lastly, Deployment DAG will handle the deployment of weights to production server.
Tasks in DAGs
For our use case of hyperparameter optimization, the prelabeling and deployment DAGs are better suited to be implemented in one node (we will refer as master node) while the training DAG is suited to be running dynamically one or more nodes (we will refer as worker nodes). This architecture minimizes complexity while still allowing the most work heavy part of our pipeline, i.e. training with different hyperparameters, to be parallelized across all the GPU devices we have. A short note regarding training parallelization we implement here is that, we want to parallelize each experiment as a whole by assigning it to a worker node rather than operator parallelization where training operations in one experiment are parallelized across several nodes. The latter form of parallelization brings another level of complexity which might not be worth dealing with in most applications.
Dashboard for recording experiment results
Tensorboard, generally, is the de-facto training dashboard for single machine training but if we want to train across different node that may also span different physical locations, using a cloud-based dashboard service might be more appropriate. Neptune.ai is one such solution I have used personally, among many other choices that are available.
Training Abstraction Layer (TAL)
Airflow is not designed to run a single task synchronously with the whole pipeline. This makes Airflow tasks unsuitable for creating a service for training tasks to communicate with to do things like abstracting away specific training dashboard, timing different parts of the training code, determining best validation and test results, managing model weights etc. To mitigate this, we can create a separate container/pod that runs a Training Abstraction Layer Service. The API calls exposed by this service can be implemented using something like XMLRPC for simplicity and each training experiment can be identified with an integer, we can call experiment_id, in every API function call parameter list.
The following are some example APIs that may be exposed by TAL:
def initialize_experiment(experiment_id: int, experiment_name: str, hyperparameters: Dict[str, Union[str, float, int]])
def end_experiment(experiment_id: int, failed: bool)
def notify_train_epoch_start(experiment_id: int, epoch: int, lr: float, _timestamp: float)
def notify_train_epoch_end(experiment_id: int, epoch: int, losses: Dict[str, float], _timestamp: float)
def notify_train_step_start(experiment_id: int, epoch: int, step: int, batch_size: int, _timestamp: float)
def notify_train_step_end(experiment_id: int, epoch: int, step: int, batch_size: int, _timestamp: float)
def notify_test_step_start(experiment_id: int, step: int, batch_size: int, _timestamp: float)
def notify_test_step_end(experiment_id: int, step: int, batch_size: int, _timestamp: float)
def notify_val_results(experiment_id: int, epoch: int, losses: Dict[str, float], metrics: Dict[str, float], _timestamp: float)
def notify_test_results(experiment_id: int, losses: Dict[str, float], metrics: Dict[str, float], _timestamp: float)
def notify_final_model_saved(experiment_id: int, weights_filename: str, _timestamp: float)
Interfacing training code with TAL and training tasks
If TAL APIs are implemented as XMLRPC server, the training code running in tasks will need to setup themselves as a XMLRPC client to call these functions. One important feature of XMLRPC is that it allows us to support function call batching. This kind of batching might be useful for API call pairs like notify_test_step_start/end listed above, as these call are usually made very fast adding cumulative network latencies to existing training latency.
One peculiar thing about the APIs defined above is that there is an extra _timestamp parameter to all the notify_ prefixed functions. This is because if we subtract timestamp between end and start of a step/epoch to time it on the server side, the calculated latency will also contain network latency with training code latency. Again, function batching will further degrade the accuracy of this timing calculation.
To solve this, we can create proxy layer code for TAL clients to use. This layer can automatically handle function batching and appending _timestamp for all notify_ calls for clients at the actual time of their call and so ensuring correctness.
Finally, we need the training code in repo to include a file containing a ‘list of hyperparameters’ so that the Airflow task responsible for creating experiments can pick one hyperparameter set from this file and dispatch a Airflow training task each. Similarly, the code in training repo should also be able to accept hyperparameter to train with using command line arguments.
Getting notification for task failure
Airflow supports setting DAG success and failure callbacks using on_success_callback and on_failure_callback. In these callbacks, we can use Airflow native operators like SlackWebhookOperator and EmailOperator to send Slack messages and email during DAG events we want to monitor.
System Architecture
The following diagram shows the full architecture of the proposed system. The master node runs the DAGs responsible for prelabeling and deployment while the worker nodes run training tasks from training DAG. Depending on the number of GPU devices available in each node, a node may take up multiple queued training tasks and run multiple trainings concurrently.
Because master and worker nodes are different machines, as shown in the figure below, they might need to share a storage mount point for reading and writing dataset and weights. To speed up training, a worker node might copy the dataset from the mount point to its local drive to avoid high latency if the mounted source is over a network. Again, when worker nodes finish their training, the mount point for weights will allow the master node to access and upload the best weight to production.
Finally, the Training Abstraction Layer (TAL), shown in the figure, runs in the master node and provides abstraction for worker node tasks as discussed in its section above.

Implementing the pipeline
Prelabeling DAG
Prelabeling DAG would consist of the following:
- A task to fetch unannotated (raw) images from warehouse storage like Amazon S3 bucket to local disk
- A task to create tables and rows for training session and information about raw images
- A task to start a labeling session with a project ID in an annotation tool like CVAT
- A task to monitor the status of the labeling session
- A task to invoke the next DAG (training DAG) with project ID of the labeling session
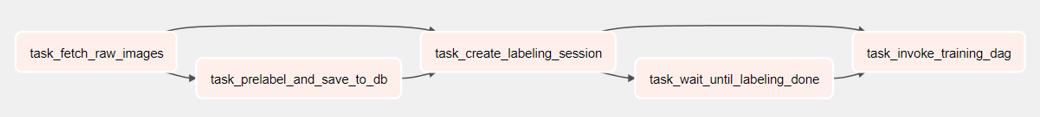
Annotation tools like CVAT report the completion status of labeling project by a status flag in their JSON response. For creating tasks that continuously pool this flag without wasting system resources (like a normal Python operator task would), we could use Airflow’s Deferred Operators. Again, running this DAG in a single node with its tasks as Celery workers should be the optimal implementation.
Deployment DAG
What deployment DAG does exactly might vary greatly depending on how a specific production environment is setup. But, generally, it would consist of the following:
- A task to report end of experiments and the best model via Slack, email etc
- A task to deploy the best model found with training experiments to production server
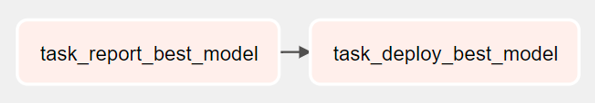
Just like in Prelabeling DAG, implementing these tasks as Celery workers seems like the most optimal method of implementation for the tasks in this DAG too. As to deploying to production server, we can either use a simple S3 bucket with custom versioning or make use of existing frameworks such as Tensorflow Serving. We probably want to manually invoke this DAG rather than automatically after Training DAG finishes.
Training DAG
The tasks in the training DAG should do the following:
- Download annotated data (identified by project ID in CVAT) from annotation tool
- Split dataset into training, validation and testing set
- Transform the dataset to the format suitable for training
- Create experiments from hyperparameters list file
- Run training and report test metrics via TAL APIs
- Determine the best model
One restriction of Apache Airflow is that tasks cannot be dynamically generated by other tasks and have to be determined at “compile” time of our DAG python script. This also means that the number of hyperparameter sets we can test must be a constant. One method of getting around this, implemented by task_check_empty_experiment in the figure for Training DAG below, is by a Python branch operator. If the number hyperparameters to test is less than the maximum number that we support (let’s say N), this branch operator can easily skip other training tasks that are not required.

Training and ranking strategy
If we just take the N experiments to test and determine the best model from them on just one shuffle of dataset using test metrices of the trained models, we run the risk of not getting the optimal best model as all the models tested have only seen that one set of dataset splits and so, we cannot be sure that the model selected this way hasn’t overfitted this set.
To mitigate this, we can do K shuffles of data and then do L experiments on each of them. This will lead to KxL number of experiments. Unfortunately, this number can get large pretty quick and hence is impractical. Rather, we can do is, first, determine N best model from M experiments with one shuffle of data. This is shown by find_best_N_models (implemented under an Airflow Task Group) in the figure above. Then, we do X epochs where we shuffle dataset to create a new set and rank the best N models found earlier, shown by find_top_model. The top model found this way will have to rank the best in three out of four experiments and hence we can be more confident about the ranking of our model.
Single machine training
If we are only going to use one machine with one GPU, running the pipeline becomes very simple. We simply run Airflow in a docker using their Docker Compose file at https://airflow.apache.org/docs/apache-airflow/<AIRFLOW_VERSION>/docker-compose.yaml. For the Airflow instance in the docker to be able to control the host docker, rather than directly changing the host machine’s file permission for /var/run/docker.sock and mapping it to the container, we can use docker service like bobrik/socat to proxy the access to the host file.
Again, as the dependency for training task i+1 is i to queue for this one GPU, they run one after another without any parallelization and need for synchronization. If there are more than one GPU and would like to queue our training tasks among them, however, we would probably want to have a similar setup as multi-machine training as discussed in the following sections.
To make sure that Airflow training task containers can access (NVIDIA) GPUs, we need to:
- Install
nvidia-docker2and set default docker runtime tonvidiaby editing/etc/docker/daemon.json. - Derive training task container images from
nvidia/cuda:<CUDA_VERSION>-cudnn<CUDNN_VERSION>-runtime-ubuntu<UBUNTU_VERSION>.
Multi-machine training using Docker Swarm
Docker Swarm allows us to build upon the single machine training setup mentioned above and create a muti-node training setup with minimal effort. We create a master node (called manager) using docker swarm init --advertise-addr <MANAGER_IP>. Then, worker nodes can notify the master node that they want to join the swarm using docker swarm join --token <TOKEN> <MANAGER_IP>. Similarly, a worker node can leave the swarm using docker swarm leave. Until Docker Compose version 3, specifying container placement (under deploy block) is not supported. So, we will have to create manager and worker node in a swarm manually. Finally, we use Airflow’s DockerSwarmOperator to put training tasks to run on worker nodes.
Problems with Docker Swarm and its support in Airflow
Unfortunately, as of the date of this post, GPU support in Docker Swarm seems immature (i.e. not easy as in Kubernetes) and Airflow’s DockerSwarmOperator doesn’t support setting GPU resource constraint via its function parameters. The former can fixed with some work by referring to https://docs.docker.com/config/containers/resource_constraints. But the latter will require actually modifying Airflow code. We need to set resource constraints so that we can assign one training task to one GPU, if a node has multiple GPUs in it. Again, there seems to be a bug in Airflow version 2.2.4 where completion status for a Docker Swarm task cannot be properly determined by Airflow if enable_logging is enabled.
(Massive) multi-machine training using Kubernetes cluster
Given the problems with Docker Swarm and its integration with Airflow, we might have to stick with Kubernetes even if we wanted to create a simple GPU-based multi-node setup. Kubernetes was a large set of components to setup and manage a multi-cluster, multi-node production environment. For our use case, we might only need few of the components as we are not creating a customer facing production environment.
Generally, Kubernetes users might use a separate third-party tools like kind, minikube etc to setup heir Kubernetes cluster. These tools provide abstraction over native Kubernetes tools like kubeadm and kubectl. Airflow internally uses these native Kubernetes tools to create, manage and delete pods for its tasks. So, if we are running Airflow in a docker, we will need to map Kubernetes config file (usually located in ~/.kube/config), network plugin CNI file (example: for flannel it would be /run/flannel/subnet.env) and install Kubernetes in the container with Airflow. This approach may not provide the best experience, however.
This is because when a Kubernetes cluster is created on the host and Docker is a separate technology to Kubernetes. Instead, we could deploy Airflow in a Kubernetes cluster’s pod running on the master machine and notify Airflow about this by setting AIRFLOW__KUBERNETES__IN_CLUSTER to true.
Next, we will need to use KubernetesPodOperator for training tasks so that Kubernetes Executor picks them up to run across worker nodes while other tasks default to Celery so that they run on master node. We also need to set the executor argument of training task as kubernetes and AIRFLOW__CELERY_KUBERNETES_EXECUTOR__KUBERNETES_QUEUE must be set to kubernetes to use Kubernetes Executor properly. Finally, to enable NVIDIA GPU support for Kubernetes we will need to install NVIDIA device plugin.
Now that we have Airflow and Kubernetes master node running, we can make worker nodes join this cluster using kubedm join <MASTER_IP> --token <TOKEN> --discovery-token-ca-cert-hash <CA_CERT_HASH>.
Queuing training tasks among available GPU resources
Besides making parallelization among nodes easier when each node has just one GPU to use, Kubernetes, using its resource constraint feature, allows us to schedule more than one task in nodes which have more than one GPU. We can make use of this feature easily using KubernetesPodOperator‘s resources function argument. We can specify the minimum number of GPUs our training code needs using data structure of type kubernetes.client.models.V1ResourceRequirements for this function argument.
Extras: Port forwarding for WSL2 setup
If you have setup your master and worker nodes with Kubernetes/Docker in WSL2, you will need to setup port forwarding from WSL VM to host using the following PowerShell script as Administrator with the following:
For Docker Swarm:
$WSL_IP_ADDRESS=$args[0]
netsh interface portproxy add v4tov4 listenport=2377 listenaddress=0.0.0.0 connectport=2377 connectaddress=$WSL_IP_ADDRESS
netsh interface portproxy add v4tov4 listenport=7946 listenaddress=0.0.0.0 connectport=7946 connectaddress=$WSL_IP_ADDRESS
netsh interface portproxy add v4tov4 listenport=4789 listenaddress=0.0.0.0 connectport=4789 connectaddress=$WSL_IP_ADDRESS
For Kubernetes:
$WSL_IP_ADDRESS=$args[0]
netsh interface portproxy add v4tov4 listenport=6443 listenaddress=0.0.0.0 connectport=6443 connectaddress=$WSL_IP_ADDRESS
netsh interface portproxy add v4tov4 listenport=2379 listenaddress=0.0.0.0 connectport=2379 connectaddress=$WSL_IP_ADDRESS
netsh interface portproxy add v4tov4 listenport=2380 listenaddress=0.0.0.0 connectport=2380 connectaddress=$WSL_IP_ADDRESS
netsh interface portproxy add v4tov4 listenport=10250 listenaddress=0.0.0.0 connectport=10250 connectaddress=$WSL_IP_ADDRESS
netsh interface portproxy add v4tov4 listenport=10259 listenaddress=0.0.0.0 connectport=10259 connectaddress=$WSL_IP_ADDRESS
netsh interface portproxy add v4tov4 listenport=10257 listenaddress=0.0.0.0 connectport=10257 connectaddress=$WSL_IP_ADDRESS



